-Search query
-Search result
Showing 1 - 50 of 488 items for (author: toma & s)

EMDB-18396: 
Cryo-EM structure of C-terminally truncated Apoptosis signal-regulating kinase 1 (ASK1)
Method: single particle / : Kosek D, Honzejkova K, Obsilova V, Obsil T

PDB-8qgy: 
Cryo-EM structure of C-terminally truncated Apoptosis signal-regulating kinase 1 (ASK1)
Method: single particle / : Kosek D, Honzejkova K, Obsilova V, Obsil T

EMDB-42601: 
CryoEM structure of Kappa Opioid Receptor bound to a semi-peptide and Gi1
Method: single particle / : Fay JF, Che T

EMDB-41071: 
Cryo-EM structure of rat cardiac sodium channel NaV1.5 with batrachotoxin analog BTX-B
Method: single particle / : Tonggu L, Wisedchaisri G, Gamal El-Din TM, Zheng N, Catterall WA

PDB-8t6l: 
Cryo-EM structure of rat cardiac sodium channel NaV1.5 with batrachotoxin analog BTX-B
Method: single particle / : Tonggu L, Wisedchaisri G, Gamal El-Din TM, Zheng N, Catterall WA

EMDB-40451: 
E1435Q Ycf1 mutant in dephosphorylated state
Method: single particle / : Khandelwal NK, Tomasiak TM

PDB-8sg4: 
E1435Q Ycf1 mutant in dephosphorylated state
Method: single particle / : Khandelwal NK, Tomasiak TM

EMDB-16229: 
Cryo-EM structure of the bacterial replication origin opening basal unwinding system
Method: single particle / : Pelliciari S, Bodet-Lefevre S, Murray H, Ilangovan A

PDB-8btg: 
Cryo-EM structure of the bacterial replication origin opening basal unwinding system
Method: single particle / : Pelliciari S, Bodet-Lefevre S, Murray H, Ilangovan A

EMDB-17757: 
Cryo-EM structure of the Cas12m-crRNA-target DNA complex
Method: single particle / : Sasnauskas G, Tamulaitiene G, Karvelis T, Bigelyte G, Siksnys V

PDB-8pm4: 
Cryo-EM structure of the Cas12m-crRNA-target DNA complex
Method: single particle / : Sasnauskas G, Tamulaitiene G, Karvelis T, Bigelyte G, Siksnys V

EMDB-19064: 
CryoEM reconstruction of SARS-CoV-2 spike in complex with nanobody tri-TMH (partially open conformation)
Method: single particle / : Rissanen I, Hannula L, Huiskonen JT

EMDB-19068: 
CryoEM reconstruction of SARS-CoV-2 Spike in complex with nanobody tri-TMH (closed conformation)
Method: single particle / : Rissanen I, Hannula L, Huiskonen JT
EMDB-34174: 
Structure of beta-arrestin2 in complex with a phosphopeptide corresponding to the human Atypical chemokine receptor 2, ACKR2 (D6R)
Method: single particle / : Maharana J, Sarma P, Yadav MK, Banerjee R, Shukla AK
EMDB-36078: 
Structure of beta-arrestin2 in complex with M2Rpp
Method: single particle / : Maharana J, Sano FK, Shihoya W, Banerjee R, Nureki O, Shukla AK
EMDB-36081: 
Structure of beta-arrestin2 in complex with D6Rpp (Local Refine)
Method: single particle / : Maharana J, Sarma P, Yadav MK, Chami M, Banerjee R, Shukla AK
EMDB-36082: 
Structure of beta-arrestin1 in complex with D6Rpp
Method: single particle / : Maharana J, Sarma P, Yadav MK, Chami M, Banerjee R, Shukla AK
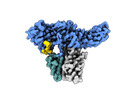
EMDB-36090: 
Structure of Muscarinic receptor (M2R) in complex with beta-arrestin1 (Local refine, cross-linked)
Method: single particle / : Maharana J, Sano FK, Shihoya W, Banerjee R, Nureki O, Shukla AK
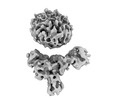
EMDB-36091: 
Muscarinic receptor (M2R) in complex with beta-arrestin1 (Low resolution full map, Cross-linked)
Method: single particle / : Maharana J, Sano FK, Shihoya W, Banerjee R, Nureki O, Shukla AK
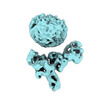
EMDB-36093: 
Muscarinic receptor (M2R) in complex with beta-arrestin1 (Low resolution full map, non-crosslinked)
Method: single particle / : Maharana J, Sano FK, Shihoya W, Banerjee R, Nureki O, Shukla AK

EMDB-36110: 
Structure of basal beta-arrestin2
Method: single particle / : Maharana J, Sarma P, Yadav MK, Chami M, Banerjee R, Shukla AK
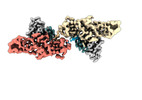
EMDB-36124: 
Structure of beta-arrestin1 in complex with C3aRpp
Method: single particle / : Maharana J, Sarma P, Yadav MK, Chami M, Banerjee R, Shukla AK
EMDB-36126: 
Structure of Muscarinic receptor (M2R) in complex with beta-arrestin1 (Local Refine, non-cross linked)
Method: single particle / : Maharana J, Sano FK, Shihoya W, Banerjee R, Nureki O, Shukla AK

PDB-8go9: 
Structure of beta-arrestin2 in complex with a phosphopeptide corresponding to the human Atypical chemokine receptor 2, ACKR2 (D6R)
Method: single particle / : Maharana J, Sarma P, Yadav MK, Banerjee R, Shukla AK

PDB-8j8r: 
Structure of beta-arrestin2 in complex with M2Rpp
Method: single particle / : Maharana J, Sano FK, Shihoya W, Banerjee R, Nureki O, Shukla AK

PDB-8j8v: 
Structure of beta-arrestin2 in complex with D6Rpp (Local Refine)
Method: single particle / : Maharana J, Sarma P, Yadav MK, Chami M, Banerjee R, Shukla AK

PDB-8j8z: 
Structure of beta-arrestin1 in complex with D6Rpp
Method: single particle / : Maharana J, Sarma P, Yadav MK, Chami M, Banerjee R, Shukla AK

PDB-8j97: 
Structure of Muscarinic receptor (M2R) in complex with beta-arrestin1 (Local refine, cross-linked)
Method: single particle / : Maharana J, Sano FK, Shihoya W, Banerjee R, Nureki O, Shukla AK

PDB-8j9k: 
Structure of basal beta-arrestin2
Method: single particle / : Maharana J, Sarma P, Yadav MK, Chami M, Banerjee R, Shukla AK

PDB-8ja3: 
Structure of beta-arrestin1 in complex with C3aRpp
Method: single particle / : Maharana J, Sarma P, Yadav MK, Chami M, Banerjee R, Shukla AK

PDB-8jaf: 
Structure of Muscarinic receptor (M2R) in complex with beta-arrestin1 (Local Refine, non-cross linked)
Method: single particle / : Maharana J, Sano FK, Shihoya W, Banerjee R, Nureki O, Shukla AK

EMDB-35029: 
SARS-CoV2 spike protein with ACE2, no ACE2 binding.
Method: single particle / : Kishikawa J, Hirose M, Kato T, Okamoto T

EMDB-35030: 
SARS-CoV2 spike protein with ACE2. 1 ACE2 bound form.
Method: single particle / : Kishikawa J, Hirose M, Kato T, Okamoto T

EMDB-35031: 
SARS-CoV2 spike protein with ACE2. 2 ACE2 bound form.
Method: single particle / : Kishikawa J, Hirose M, Kato T, Okamoto T

EMDB-35032: 
SARS-CoV2 spike protein with ACE2. 3 ACE2 bound form.
Method: single particle / : Kishikawa J, Hirose M, Kato T, Okamoto T

EMDB-35036: 
SARS-CoV2 spike protein with ACE2 decoy.no ACE2 decoy binding
Method: single particle / : Kishikawa J, Hirose M, Kato T, Okamoto T

EMDB-35037: 
SARS-CoV2 spike protein with ACE2 decoy. 1 ACE2 decoy bound form.
Method: single particle / : Kishikawa J, Hirose M, Kato T, Okamoto T

EMDB-35038: 
SARS-CoV2 spike protein with ACE2 decoy. 1 ACE2 decoy bound and 2 RBD up form.
Method: single particle / : Kishikawa J, Hirose M, Kato T, Okamoto T

EMDB-35039: 
SARS-CoV2 spike protein with ACE2 decoy. 2 ACE2 decoy bound form.
Method: single particle / : Kishikawa J, Hirose M, Kato T, Okamoto T

EMDB-35040: 
SARS-CoV2 spike protein with ACE2 decoy. 3 ACE2 decoy bound form.
Method: single particle / : Kishikawa J, Hirose M, Kato T, Okamoto T

EMDB-36345: 
RBD of SARS-CoV2 spike protein with ACE2 decoy
Method: single particle / : Kishikawa J, Hirose M, Kato T, Okamoto T

PDB-8jje: 
RBD of SARS-CoV2 spike protein with ACE2 decoy
Method: single particle / : Kishikawa J, Hirose M, Kato T, Okamoto T

EMDB-29026: 
CryoEM structure of Kappa Opioid Receptor bound to a semi-peptide and Gi1
Method: single particle / : Fay JF, Che T

PDB-8feg: 
CryoEM structure of Kappa Opioid Receptor bound to a semi-peptide and Gi1
Method: single particle / : Fay JF, Che T

EMDB-16005: 
GABA-A receptor a5 homomer - a5V3 - APO
Method: single particle / : Miller PS, Malinauskas TM, Hardwick SW, Chirgadze DY

EMDB-16050: 
GABA-A receptor a5 homomer - a5V3 - Basmisanil - HR
Method: single particle / : Malinauskas TM, Wahid AA, Hardwick SW, Chirgadze DY, Miller PS

EMDB-16051: 
GABA-A receptor a5 homomer - a5V3 - RO154513
Method: single particle / : Miller PS, Malinauskas TM, Hardwick SW, Chirgadze DY, Wahid AA

EMDB-16055: 
GABA-A receptor a5 homomer - a5V3 - RO5211223
Method: single particle / : Miller PS, Malinauskas TM, Hardwick SW, Chirgadze DY

EMDB-16058: 
GABA-A receptor a5 homomer - a5V3 - Diazepam
Method: single particle / : Miller PS, Malinauskas TM, Kasaragod VB, Chirgadze DY

EMDB-16060: 
GABA-A receptor a5 homomer - a5V3 - DMCM
Method: single particle / : Miller PS, Malinauskas TM, Hardwick SW, Chirgadze DY
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
